Dynamical Domain Decomposition¶
Let’s start with essential import statements:
In [1]: from prody import *
In [2]: import numpy as np
We’ll start by parsing the pseudo-atoms from a PDB file generated in the previous step.
In [3]: emd = parsePDB('EMD-1961.pdb')
The order of pseudo-atoms generated by TRN is random and does not follow a sequence like residues in an atomic model do. Nor do they have no chain identifiers.
As an alternative to reordering the pseudo-atoms and assigning residue and chain identifers
to the pseudo-atoms based on the PDB structure in the previous section, we demonstrate here
the use of dynamical domain decomposition using GNM modes with the function calcGNMDomains().
The number of dynamical domains obtained is approximately equal to the number of modes use so
we use 8 modes to get about 8 domains as in our recent paper ([YZ20]).
In [4]: gnm, _ = calcGNM(emd, selstr='all')
In [5]: domains = calcGNMDomains(gnm[:8])
In [6]: print(np.unique(domains))
[0 1 2 3 4 5 6 7 8]
We actually get 9 in this case as you will see in the figure at the end.
We can assign this data to the AtomGroup as follows:
In [7]: emd.setData('domain', domains)
This allows the domains to be used for selection:
In [8]: domain1 = emd.select('domain 1')
In [9]: print(domain1.numAtoms())
199
Finally, we save a domain-labelled pseudo-atom model to a PDB file for visualization and other downstream analyses, using the B-factor field for writing the domains:
In [10]: writePDB('EMD-1961_domains.pdb', emd, beta=domains)
Out[10]: 'EMD-1961_domains.pdb'
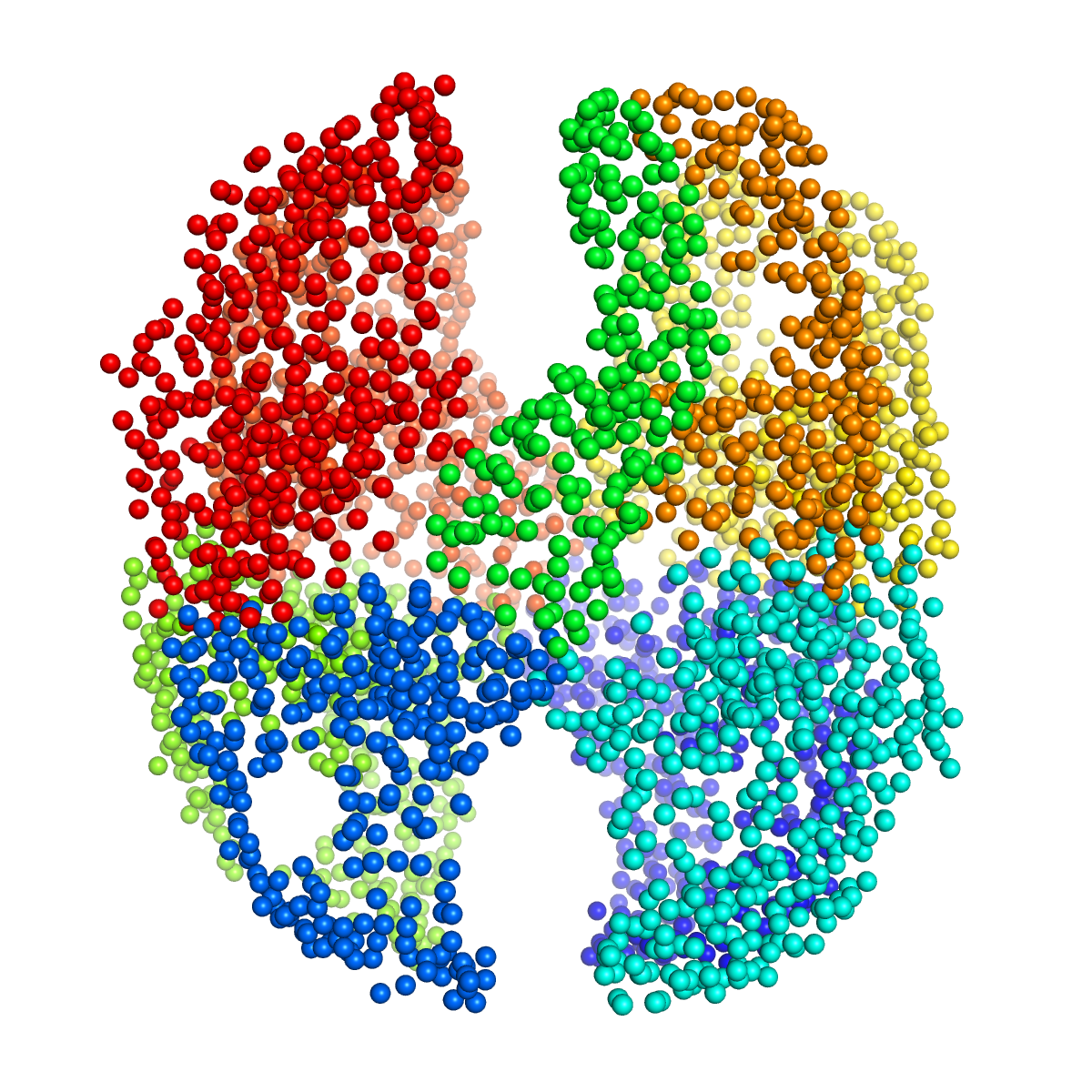
Visualisation of the resultant structure will look something like the following figure.
We see 9 domains, including 7 pairs of domains and 2 individual domains.
Another round of decomposition with each of the resulting domains can be used to divide them further. For example, we can decompose the first domain (index 0) into 4 dynamical domains using the first 3 modes, which can be combined to create subunits or used as they are for comparison with dynamical domains for atomic models.
In [11]: gnm0, domain0 = calcGNM(emd, selstr='domain 0')
In [12]: domains0 = calcGNMDomains(gnm0[:3])
In [13]: writePDB('EMD-1961_domain0_subdomains4.pdb', domain0, beta=domains0)
Out[13]: 'EMD-1961_domain0_subdomains4.pdb'