Explicit Membrane ANM¶
As in the previous section, we will investigate the effect of the presence of membrane for the motion of a neurotransmitter transporter with ProDy’s explicit membrane ANM (exANM) capabilities. The procedure is based on the methods described in [TL12] and relies on building a real membrane and using reduceModel().
Tutorial Files¶
Files in the following archives can be used to follow this tutorial:
Here is a list of these files:
7.5K May 10 09:33 2nwl_blocks.txt
1008K May 10 09:33 2NWL-opm.pdb
7.6K May 10 09:33 3kbc_blocks.txt
1.1M May 10 09:33 3KBC-opm.pdb
The file contains the outward-facing structure of the glutamate transporter after insertion into the plasma membrane. It is obtained from the Orientations of Proteins in Membranes database.
| [TL12] | Lezon TR, Bahar I. Constraints Imposed by the Membrane Selectively Guide the Alternating Access Dynamics of the Glutamate Transporter GltPh. Biophys J 2012 102 1331-1340. |
Preparing the structures¶
We recommend that you will follow this tutorial by typing commands in an IPython session, e.g.:
$ ipython
or with pylab environment:
$ ipython --pylab
First, we will make necessary imports from ProDy and Matplotlib packages.
In [1]: from prody import *
In [2]: from pylab import *
In [3]: ion()
The exANM assumes that the membrane is normal to the z-axis, so it is important to use a structure that is properly aligned. The structure from the OPM database will work.
In [4]: of_all = parsePDB('2NWL-opm.pdb') # Outward-facing structure
There will be warnings saying that ProDy wants to read beta factors, but the coordinates should be read properly. In addition to atoms, the OPM file contains points to indicate the boundaries of the membrane.
Building Hessian and Calculating Modes¶
ProDy’s exANM method can be used for any system. This method will create a membrane with given highest and lowest coordinate on the Z-axis. The main advantage of this method is that the protein can interact with lipid molecules on the membrane. The elastic network model based on the interaction between aminoacids on protein and the interaction between aminoacids with the lipids on membrane. The addition of physical membrane will avoid the unphysical distortion of the structure. This will not reduce accuracy as in the case of implicit membrane ANM model.
To use the explicit membrane for ANM calculation, we first select the protein part of the structure:
In [5]: of_ca = of_all.select('protein and name CA and not (chain A and resid 119 to 122) and not (chain C and resid 119 to 123) and not chain D')
and then instantiate an exANM object:
In [6]: exanm = exANM('2nwl')
and we build a couple of Hessians using the coordinates of the crystal structures,
In [7]: exanm.buildHessian(of_ca, h=13.)
h=13. specifies the thickness of the membrane to be 26 Angstrom. Now we calculate the modes and write them to a pair of .nmd files for viewing.
In [8]: exanm.calcModes()
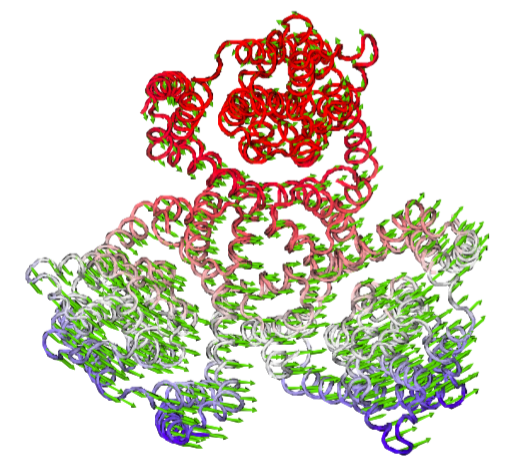
In [9]: writeNMD('2nwl_ex.nmd',exanm,of_ca.select('protein and name CA'))
Out[9]: '2nwl_ex.nmd'