Structure Analysis¶
ProDy comes with many functions that can be used to calculate structural properties and compare structures. We demonstrate only some of these functions. For more detailed examples, see Structure Analysis tutorial.
In [1]: from prody import *
In [2]: from pylab import *
In [3]: ion()
Measure geometric properties¶
Let’s parse a structure:
In [4]: p38 = parsePDB('1p38')
Functions for analyzing structures can be found in measure
module. For example, you can calculate phi (φ) and psi (ψ) for the 10th
residue, or the radius of gyration of the protein as follows:
In [5]: calcPhi(p38[10,])
Out[5]: -115.5351427673999
In [6]: calcPsi(p38[10,])
Out[6]: 147.49025666398765
In [7]: calcGyradius(p38)
Out[7]: 22.057752024921772
Compare and align structures¶
You can also compare different structures using some of the methods in
proteins module. Let’s parse another p38 MAP kinase structure
In [8]: bound = parsePDB('1zz2')
You can find similar chains in structure 1p38 and 1zz2 using
matchChains() function:
In [9]: apo_chA, bnd_chA, seqid, overlap = matchChains(p38, bound)[0]
In [10]: apo_chA
Out[10]: <AtomMap: Chain A from 1p38 -> Chain A from 1zz2 from 1p38 (337 atoms)>
In [11]: bnd_chA
Out[11]: <AtomMap: Chain A from 1zz2 -> Chain A from 1p38 from 1zz2 (337 atoms)>
In [12]: seqid
Out[12]: 99.40652818991099
In [13]: overlap
Out[13]: 96
Matching Cα atoms are selected and returned as AtomMap instances.
We can use them to calculate RMSD and superpose structures.
In [14]: calcRMSD(bnd_chA, apo_chA)
Out[14]: 72.93023086946586
In [15]: bnd_chA, transformation = superpose(bnd_chA, apo_chA)
In [16]: calcRMSD(bnd_chA, apo_chA)
Out[16]: 1.8628014908695476
In [17]: showProtein(p38);
In [18]: showProtein(bound);
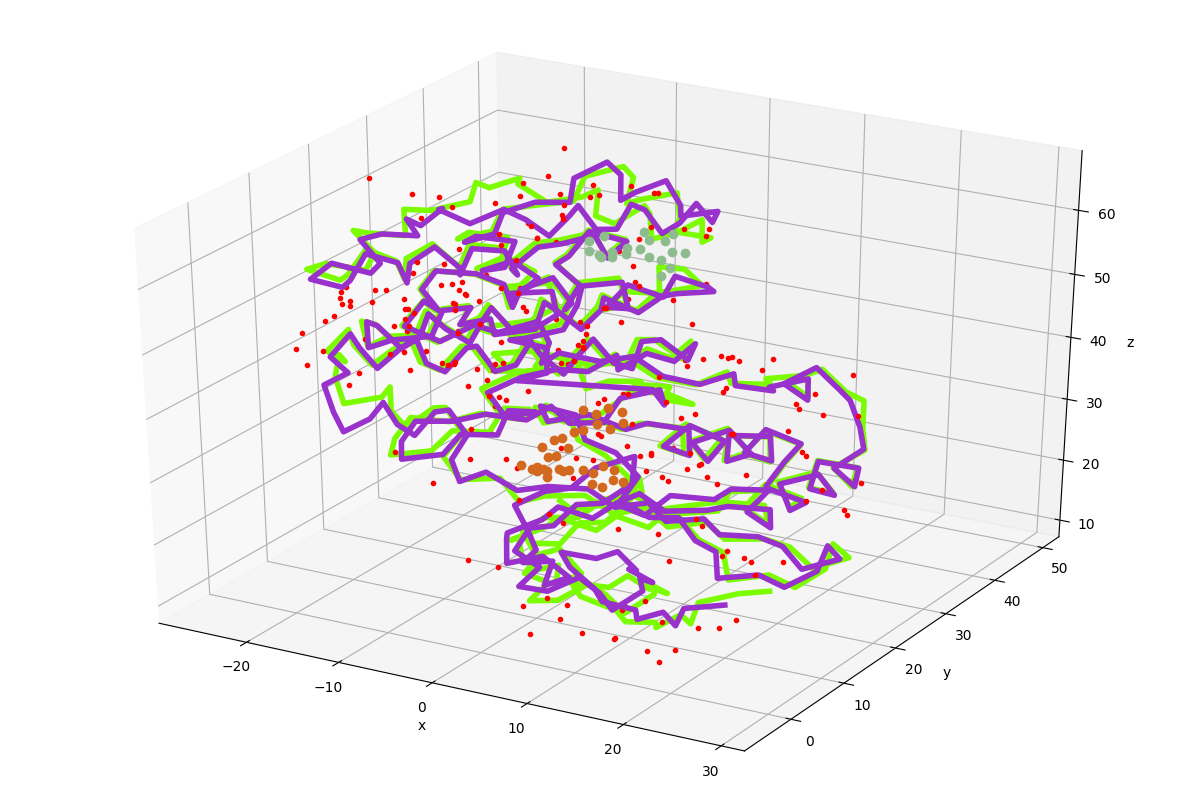
Writing PDB files¶
PDB files can be written using the writePDB() function.
The function accepts objects containing or referring to atomic data.
Output selected atoms:
In [19]: writePDB('1p38_calphas.pdb', p38.select('calpha'))
Out[19]: '1p38_calphas.pdb'
Output a chain:
In [20]: chain_A = p38['A']
In [21]: writePDB('1p38_chain_A.pdb', chain_A)
Out[21]: '1p38_chain_A.pdb'
As you may have noticed, this function returns the file name after it is successfully written. This is a general behavior for ProDy output functions. For more PDB writing examples see Write PDB file.
